missing translation for 'onlineSavingsMsg'
Learn More
Learn More
Invitrogen™ TREX1 Polyclonal Antibody


Description
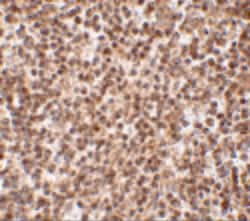
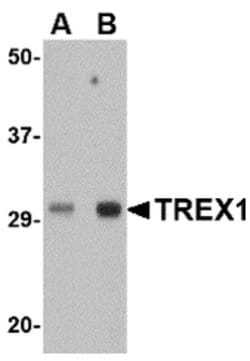
A suggested positive control is human spleen tissue lysate. PA5-20665 can be used with blocking peptide PEP-0785.
Trex1 (ATRIP) is the major human 3' to 5' exonuclease which is required for checkpoint signaling after DNA damage. It is ubiquitously expressed, binds to single stranded DNA coated with replication protein A that accumulates at sites of DNA damage and recruits the ataxia telangiectasia and Rad3 related protein (ATR), a checkpoint kinase, to sites of DNA damage and replication stress. Trex1 is required for ATR expression. This gene uses two different open reading frames. The upstream ORF encodes proteins which interact with ATR and localize to intranuclear foci induced by DNA damage and are essential components of the DNA damage checkpoint. The downstream ORF encodes proteins with 3' to 5' exonuclease activity and may be a subunit of human DNA polymerase III. Multiple transcript variants encoding different isoforms have been found. Mutations in this gene result in Aicardi-Goutieres syndrome, chilblain lupus, and Cree encephalitis.

Specifications
Specifications
| Antigen | TREX1 |
| Applications | Immunohistochemistry, Western Blot |
| Classification | Polyclonal |
| Concentration | 1 mg/mL |
| Conjugate | Unconjugated |
| Formulation | PBS with 0.02% sodium azide |
| Gene | TREX1 |
| Gene Accession No. | Q91XB0, Q9NSU2 |
| Gene Alias | 1661; 3' repair exonuclease 1; 3'-5' exonuclease TREX1; AGS1; AU041952; CRV; Deoxyribonuclease III; DNase III; DRN3; HERNS; RGD1309596; three prime exonuclease 1; three prime repair exonuclease 1; three-prime repair exonuclease 1; Trex1; trophoblast expressed 1 |
| Gene Symbols | TREX1 |
| Show More |
Product Title
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.
Spot an opportunity for improvement?